- Search NCIBI Data
(e.g. diabetes, csf1r) - Login

 SAGA / TALE
SAGA / TALE
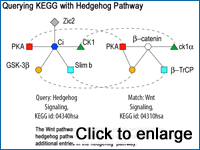
Matches Cytoscape sub-graphs to KEGG pathways.
SAGA (Substructure Index-based Approximate Graph Alignment) is a tool for querying a biological graph database to retrieve matches between subgraphs of molecular interactions and biological networks. SAGA implements an efficient approximate subgraph matching algorithm that can be used for a variety of biological graph matching problems such as the pathway matching SAGA uses to compare pathways in KEGG and Reactome.
You can also use SAGA to find matches in literature databases that have been parsed into semantic graphs. In this use of SAGA, portions of PubMed have been parsed into graphs that have nodes representing gene names. A link is drawn between two genes if they are discussed in the same sentence (indicating there is potential association between the two genes). SAGA lets you match graphs between different databases even though the content is distinct and the databases organize pathways in different ways. This cross-database matching is achieved by SAGAs flexible approximate subgraph matching model that computes graph similarity, and allows for node gaps, node mismatches, and graph structural differences. Comparing pathways from different databases can be a useful precursor to pathway data integration.
SAGA is very efficient for querying relatively small graphs, but becomes prohibitory expensive for querying large graphs. Large graph data sets are common in many emerging database applications, and most notably in large-scale scientific applications. To fully exploit the wealth of information encoded in graphs, efective and efficient graph matching tools are critical. Due to the noisy and incomplete nature of real graph datasets, approximate, rather than exact, graph matching is required. Furthermore, many modern applications need to query large graphs, each of which has hundreds to thousands of nodes and edges. TALE is an approximate subgraph matching tool for matching graph queries with a large number of nodes and edges. TALE employs a novel indexing technique that achieves a high pruning power and scales linearly with the database size.
References

 NCIBI on Facebook
NCIBI on Facebook NCIBI RSS Feed
NCIBI RSS Feed

